Select Projects
Modeling invasive species short- and long-term population dynamics to predict invasion successShort-term transient population dynamics are common in vertebrates, particularly invasive vertebrates, and by their nature are directly influenced by the interaction of population structure and vital rates. Using a novel methodological framework, we found consistent differences in the way vital rates and age structure in invasive and native populations contribute to transient dynamics suggesting that invasive and native populations are influenced by differing mechanisms. These dynamics appear to be linked with environmental conditions that regulate demography. Vital rates with the largest influence on population growth had the greatest variability across populations, contrary to the demographic buffering hypothesis. Our findings indicate the potential of a new hypothesis that could be generalized for invasive species, suggesting that tradeoffs in sensitivity between vital rates and age structure can buffer populations. We also found surprising potential for strong contributions of juvenile classes and a smaller direct role for fecundity than previously thought. While relevant to all species, these insights are particularly useful for understanding the response of invasive species and hence developing appropriate, ecologically informed control methods using “best science” practices. This work is on-going and currently being expanded to further explore the sensitivity of lower level vital rates.
|
Determining transmission potential for pathogens between wildlife and livestock
In the last half century, significant attention has been given to animal diseases; however, our understanding of disease processes and how to manage them at the livestock–wildlife interface remains limited. In these studies, we conduct a systematic review of the scientific literature to evaluate the status of diseases at the livestock–wildlife interface. We use transmission potential networks to better understand risk of transmission. Specifically, the goals of these meta-analyses were to evaluate the presence of animal diseases that can be transmitted at the wildlife-livestock interface; second to identify critical issues faced in managing these diseases at the livestock–wildlife interface; and third to identify potential technical and policy strategies for addressing these issues. These studies have found that a large percentage of pathogens reportable to the World Organization for Animal Health (OIE) can be transmitted between livestock and wildlife. The complex nature of these systems highlights the need to understand the role of wildlife in the epidemiology, transmission, and maintenance of infectious diseases of livestock. Successful management or eradication of these diseases will require the development of cross-discipline and institutional collaborations. Despite social and policy challenges, there remain opportunities to develop new collaborations and new technologies to mitigate the risks posed at the livestock–wildlife interface.
|
Modeling pathogen prevalence and disease dynamics in wildlife populations
Studies across a range of host-pathogen systems indicate that host species’ diversity can regulate the ability for a pathogen to establish. Studies of natural populations have examined correlations between the spatial distribution of disease and environmental variables, particularly those that can encourage immunologic susceptibility in a population. However there may be trade-offs between species diversity and environmental conditions, and this may be particularly important for invasive species or species pioneering new range. The interaction of environmental conditions influencing the host or pathogen may operate at scales different than those of species diversity generating asymmetric effects on pathogen transmission. These effects may also be different for pathogens with a narrow versus large host range. Yet studies investigating relationships between pathogen persistence, environmental factors, and species diversity for pathogens with narrow and wide host ranges are still relatively limited, particularly , particularly at the macro-scale and for mammal species in North America. These studies use hierarchical Bayesian models that accounts for imperfect detection probability to investigate the influence of environmental conditions, species diversity, and demographics on the infection probability in wild pigs for pathogens with broad and narrow host ranges.
|
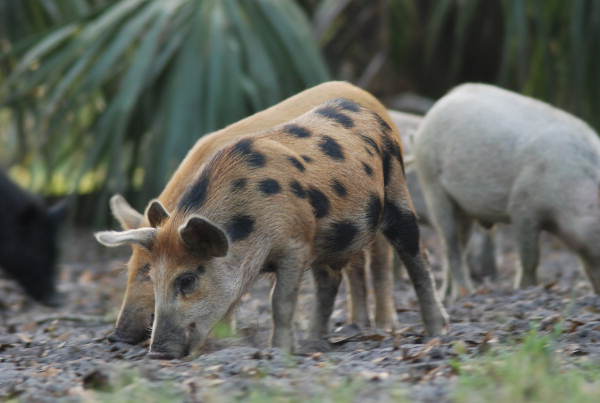
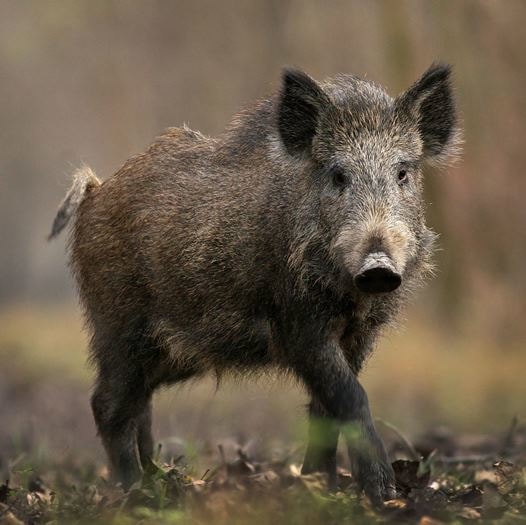
Invasive species range expansion and population dynamics
In the United States the annual damage and control costs associated with invasive invertebrates is estimated to be at least $46 billion annually. Feral swine are the most abundant free-ranging, exotic ungulate in the United States. Since the 1960s feral swine have expanded their range to at least 38 States and 3 provinces in Canada. Feral swine populations continue to increase due in part to their reproductive capacity, adaptable biology, lack of natural predators, and intentional or accidental introduction by humans. This range expansion and ability to readily adapt and thrive in a diversity of habitats has contributed to the impact of feral swine on ecosystems and agricultural systems in North America. These models use a diversity of approaches that range from Hierarchical Bayesian approaches to mathematical population models to predict the invasion of wild pigs into North America and identify parameters predictive of invasion success.
|
Modeling the impact of animal movements on transmission dynamics of disease at local and national scales.
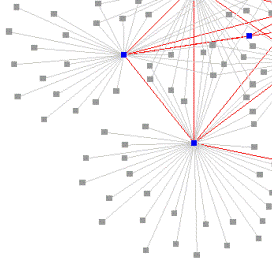
Globalization has increased the potential for the introduction and spread of novel pathogens over large spatial scales necessitating continental-scale disease models to guide emergency preparedness. Livestock disease spread models, such as those for the 2001 foot-and-mouth disease (FMD) epidemic in the United Kingdom, represent some of the best case studies of large-scale disease spread. However, generalization of these models to explore disease outcomes in other systems, such as the United States’s cattle industry, has been hampered by differences in system size and complexity and the absence of suitable livestock movement data. These studies focus on improving our understanding of the role animal movement plays in disease spread at local and national scales. Results from these studies highlight the role of large-scale disease models in emergency preparedness, particularly for systems lacking comprehensive movement and outbreak data, and the need to rapidly implement multi-scale contingency plans during outbreaks.
|
Transmission of Bovine Tuberculosis (Mycobacterium bovis) between wildlife and cattle
Mycobacterium bovis (M. bovis), the causative agent of bovine tuberculosis, has been identified in nine geographically distinct wildlife populations in North America and Hawaii and is endemic in at least three populations, including members of the Bovidae, Cervidae, and Suidae families. The emergence of M. bovis in North American wildlife poses a serious and growing risk for livestock and human health and for the recreational hunting industry. Experience in many countries, including the USA and Canada, has shown that while M. bovis can be controlled when restricted to livestock species, it is almost impossible to eradicate once it has spread into ecosystems with free-ranging maintenance hosts. Therefore, preventing transmission of M. bovis to wildlife may be the most effective way to mitigate economic and health costs of this bacterial pathogen. The objective of these projects is to better understand the status of M. bovis infection in wildlife, the risk factors important for transmission, and mitigation strategies.
|
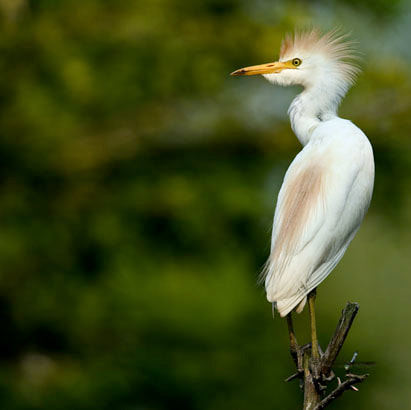
Assessment of Risk for Introduction of Heartwater (Ehrlichia ruminantium) into the Continental United States by Cattle Egrets.
Heartwater, caused by Ehrlichia (formerly Cowdria) ruminantium, is an infectious but non-contagious tick-borne disease of domestic and wild ruminants that often results in death of clinically affected individuals. This disease was first described in South Africa in 1838 as a nervous condition of sheep following a massive infestation of ticks. In 2008, many subsaharan Africa countries and Guadeloupe, in the Caribbean islands, reported clinical heartwater in their domestic ruminant livestock populations. In years past, this disease has also killed livestock on Marie-Galante and Antigua in the Caribbean.
Given the proximity of the U.S. mainland to the Caribbean islands, the ultimate concern for animal health officials, the livestock industry, and other stakeholders may not be what should be done to keep heartwater from being introduced into the continental U.S., but rather what can be done to limit the extent of an outbreak when this incursion occurs. A key element to controlling the extent of a disease outbreak is early recognition of its presence in an animal population. Consequently, the objective of this project is to determine the potential risk for introduction for of Ehrlichia ruminantium (Heartwater) into the United States mainland. A secondary goal of the project is to present the salient features of heartwater to bring increased disease awareness to veterinarians, livestock owners, wildlife biologists, and others. |
Determining differences in the spatial distribution of forest structure on the Kaibab Plateau: Implications for forest management and the northern goshawk.
The Kaibab Plateau, in North Central Arizona, has undergone extensive change in the last 100 years due to land management practices such as logging, road building, and fire suppression. The northern goshawk (Accipiter gentilis), a sensitive species, has been a center of controversy, due to the potential effects of silvicultural practices on goshawk breeding habitat. Consequently, landscape scale management plans have been proposed for goshawks in both the Pacific Northwest and Desert Southwest. This study investigated possible forest structure differences between the North Kaibab Ranger District and the adjacent Grand Canyon National Park. Forest inventory data was collected for both the Grand Canyon National Park and North Kaibab Ranger District. Analysis was conducted at three scales biologically important to the northern goshawk: landscape, stand, and nest site.
|
Farm Location and Animal Population Simulator: A system for estimating farm and animal populations.
A significant limitation in developing accurate and meaningful animal disease simulation models in the United States is the lack of data on locations of individual farms and on the numbers and types of animals on each farm. However, the United States National Agricultural Statistics Service (NASS) Census of Agriculture collects and provides data on numbers of farms and animals by county and postal zip code. In most cases privacy rules and legislation prevent the distribution of information on individual farms to most federal agencies or the public by NASS.
The Farm Location and Animal Population Simulator generates agriculturally and geographically realistic datasets based on the NASS data for disease simulation models. The simulator uses NASS data to estimate the number and sizes of animal facilities within a geographic region (e.g., states, or counties). Over the region of interest, geographic constraints are applied so that facilities are not located in urban areas, in lakes, parks, natural areas, or other locations where they are unlikely to occur. Facilities are located randomly, unless a weight matrix is used to concentrate certain types of farms closer to urban areas, live animal markets, and feed suppliers. Numbers of animals are assigned using a distribution. This simulator is currently being validated with true farm location data for North Carolina and Arkansas. Validation of the simulator fill be two fold; first the spatial distribution and assignment of animal populations on those farms will be assessed to determine if the patterns are similar to the true distribution of farms and associated populations. Second, disease spread models will be run on both the true data and the simulated data to determine how simulated population data affects the results of disease spread models. This project is currently on going. |
Linking the Pesticide Use Reporting Data with Spatial Land Use Data for Exposure Assessment
In regions of intense agricultural production, the potential for exposure to agricultural pesticides has become an important topic of public health concern. Pesticide transport into watersheds and potential affects on amphibian populations has also become a concern for environmental scientists. The State of California has developed a Pesticide Use Reporting Database (CPUR), with the objective of providing complete pesticide-use data for evaluating possible associations with human illness clusters and potential environmental affects. However, the reporting unit for the database is 1 mi2, which may be too large for accurately predicting exposure at the individual residence level necessary for some epidemiological studies of wildlife and humans.
We used the California Department of Water Resources (CDWR) crop map database to improve the crop location attributes of the CPUR database. We generated exposure metrics based on CPUR alone and CPUR linked to CDWR for birth residences in 1988-1994 in a childhood cancer study conducted by the California Department for Health Services. Sixty-six residences had both CPUR and CDWR data for the child's birth year. We calculated metrics predicting the lbs/mile2 of pesticide for 6 pesticides with high use in the study area: herbicides, simazine and trifluralin; insecticides, dicofol and propargite; and fumigants, methyl bromide and metam sodium. We first compared the exposure classification (exposed / not exposed) to each pesticide and evaluated agreement using a chi-squared test and Cohen's kappa. We then assessed differences in predicted lbs/mile2 of pesticide within a 500-meter buffer around each residence between the metrics, a distance used in previous studies of pesticide exposure. Four of the six pesticides, simazine, trifluralin, dicofol, and methyle bromide, indicated similar categorical assignment of exposure for each metric. Cohen's kappa values for these pesticides ranged from 0.11 to 0.66. The Wilcoxon signed-rank test indicated significant differences in estimated lbs/mile2 applied within the residential buffers between the CDWR metric and the CPUR metric for 3 of the pesticides. The medians and interquartile ranges were propargite, CDWR = 0.04 (0.00-0.57) and CPUR = 89.41 (14.47-233.69); simazine, CDWR = 0.00 (0.00-0.03) and CPUR = 8.77 (0.00-50.13); methyl bromide, CDWR = 0.00 (0.00-0.00) and CPUR = 0.00 (0.00-342.27). These pesticide exposure metrics are pending field validation but show promise in predicting potential pesticide exposure. Exposure metrics that refine the locational attributes of pesticide use data such as CPUR may reduce exposure misclassification for subjects (wildlife or human) with high or low exposure. |
Assessing the Spatial Accuracy of the Pesticide Use Reporting Data for Use in Exposure Assessment Studies
Exposure to agricultural chemicals has been associated with disease outcomes such as cancer, immune system disorders, adverse reproductive outcomes, developmental disorders, and neurological disease in both humans and animals. In addition, there is evidence that pesticides may severely affect amphibian reproduction and development. Pesticide use data that provides accurate location for where pesticides are applied is important for assessing relationships between disease and pesticide exposure. The State of California has developed a Pesticide Use Reporting Database (CPUR), with the objective of providing complete agricultural pesticide use data for evaluating possible associations with disease in both humans and animals. Several recent epidemiological investigations have utilized the CPUR database. However, the spatial accuracy of the database has not been evaluated. The CPUR database was established in the 1950's with limited reporting and contains information on the pesticide type, pounds applied, date, method of application, acres of crop treated. Beginning in 1990, a full use reporting system was instituted requiring commercial growers to report all pesticides used in agriculture. The reporting unit for the database is Public Land Survey System Section, which is approximately 1 mi2.
We compared the CPUR pesticide application data by crop with high precision, 100% ground verified land-use data collected by the California Department of Water Resources (CDWR). CDWR identifies 83 specific crop types with a minimum mapping unit of 0.003 mi2. We used a Geographic Information System (GIS), to conduct a comparative analysis of the location of pesticide application by crop, as reported in CPUR, with the location of the same crops in the CDWR database. We conducted the comparative analysis for ten crops with the greatest number of pesticide applications for two counties, San Joaquin (1988) and Kings (1991). To assess the accuracy of the CPUR data, we developed a GIS analytical procedure, which computed the spatial agreement between the two datasets. The comparative analysis was performed at the Section extent (N = 3,856). We tested the resulting congruence estimates for statistical significance using a one-sided binomial test for two levels of assumed crop location error in the CDWR database (1% and 5% error). Overall agreement between CPUR and CDWR was relatively high, although statistically significant differences existed between the datasets assuming 5% error in CDWR data. Accuracy assessment indicates large variation in CPUR reporting accuracy, ranging from 73.1% for cherries to 95.1% for cotton. In general, overall agreement was significantly higher in Kings County for 1991 than for San Joaquin County in 1988 for both non-permanent crops (92.9% vs 81.9% respectively) and semi-permanent crops (87.4% vs 80.7% respectively). These results indicate spatial and temporal differences in accuracy of the CPUR dataset, at both the aggregate and individual crop level. The spatial accuracy of pesticide-use data can affect exposure assessment, and may result in misclassification of exposure in epidemiological investigations of wildlife (amphibians) or humans. |
Identifying Populations Potentially Exposed to Agricultural Pesticides Using Remote Sensing and a Geographic Information System
Pesticides used in agriculture may cause adverse health effects among human and wildlife populations living near agricultural areas. However, identifying populations most likely to be exposed is difficult. We conducted a feasibility study to determine whether satellite imagery could be used to reconstruct historical crop patterns. We used historical Farm Service Agency records as a source of ground reference data to classify a late summer 1984 satellite image into crop species in a three-county area in south central Nebraska. Residences from a population-based epidemiologic study of non-Hodgkin lymphoma were located on the crop maps using a geographic information system (GIS). Corn, soybeans, sorghum, and alfalfa were the major crops grown in the study area.
Eighty-five percent of residences could be located, and of these 22% had one of the four major crops within 500 meters of the residence, an intermediate distance for the range of drift effects from pesticides applied in agriculture. We determined the proximity of residences to specific crop species and calculated crop-specific probabilities of pesticide use based on available data. This feasibility study demonstrated that remote sensing data and historical records on crop location can be used to create historical crop maps. The crop pesticides that were likely to have been applied can be estimated when information about crop-specific pesticide use is available. Using a GIS, zones of potential exposure to agricultural pesticides and proximity measures can be determined for residences in a study. |